Electron Microscopy - Tecnai G2 TEM

PPMS System Name: Tecnai G2 TEM
Department and Room: Physiology, Development and Neuroscience, Anatomy Building, Room 20
Voltages: 80, 120 and 200 keV
Emission source: LaB6
Stage: compu-stage with ± 60 degree stage tilt for 3D tomography
Detectors/Cameras: bottom-mounted AMT CCD camera
Capabilities:
- High angle annular darkfield STEM (HAADF-STEM) for Z-contrast imaging.
- Elemental analysis by EDX (Peltier-cooled Ametek silicone drift detector; Genesis/ Team software)
- Two single tilt holders, one low background EDX holder, one tomography holder
Back to main Electron Microscopy page
Bright-field TEM of Resin Sections
Biological samples typically require extensive preparation for bright-field TEM, as they contain high water content and low atomic number elements, resulting in poor inherent contrast.
Standard preparation includes:
- Fixation – to preserve cellular structure in a near-native state.
- Heavy metal staining – using agents such as osmium tetroxide, uranyl acetate, and lead citrate to enhance contrast.
- Dehydration and resin embedding – to enable ultrathin sectioning for TEM imaging.
There is no universal protocol, as methods must be tailored to the sample type. This is particularly important when preserving inorganic inclusions (e.g. nanoparticles) for EDX or EELS analysis, or when preparing samples for immuno-TEM, where maintaining antigenicity is critical.

Culture of M3T3 cells induced in-vitro to produce extracellular matrix (ECM) and matrix calcifications (C). Low-magnification overview image; the knife mark striations are caused by the inherent calcium phosphate crystals (doi.org/10.1016/j.celrep.2019.05.038). Rakesh Rajan, Prof. Melinda Duer Group, Department of Chemistry, Cambridge

Uptake of multi-walled carbon nanotubes by human macrophages in culture; arrowheads indicate MWCNTs, m = mitochondria, v = intracellular vesicles; scale bar is 100 nm (doi: 10.1021/acsbiomaterials.6b00197). Prof. Slawomir Boncel, Silesian University of Technology, Poland

Low magnification overview of a macrophage taking up vast amounts of HA nanoparticles. The particles are sequestered in large, interconnected vacuoles (arrows) that form a surface-connected compartment; n = nucleus. (doi.org/10.1016/j.biomaterials.2013.10.041). Karin Müller, Prof. Melinda Duer, Yusuf Hamied Dept. of Chemistry, Cambridge
Unstained samples
Some specimens composed of elements with higher atomic numbers are inherently electron-dense and can be imaged without applying any staining. Many crystalline powders or engineered nanoparticles for instance can simply be applied to lacey or holey carbon film grids and can immediately be imaged in bright-field TEM or STEM mode, or their elemental composition and crystallinity can be analysed by (Energy-Dispersive X-ray Spectroscopy) EDX and Selected Area Electron Diffraction (SAED).

Calcium phosphate wet precipitate before (left) and after (right) autoclave treatment (Karin Müller, Microscopy Bioscience Platform-EM, doi.org/10.1016/j.biomaterials.2013.10.041).

PAR-Ca spheres formed after incubation of the polysaccharide poly (ADP ribose) with calcium ions. The EDX spectrum shows peaks for PAR-bound calcium, the phosphorus peak is derived from the pyrophosphate groups of PAR. The SAED shows the amorphous nature of these spheres (Karin Müller; Prof. Melinda Duer – Dept. of Chemistry, Cambridge; Prof. Cathy Shanahan – King’s College London).
Negative Staining
Negative staining is a rapid and straightforward method for preparing samples for TEM. Electron-dense stains such as uranyl acetate, ammonium molybdate, or phosphotungstic acid are used to provide contrast for specimens adsorbed onto TEM grids. This technique is well-suited for visualising larger proteins, aggregates, filaments, membrane vesicles, bacteria, and viruses. It can also be combined with immunogold labelling, using a primary antibody and a gold-tagged secondary antibody, to reveal specimen morphology and detect specific surface antigens.
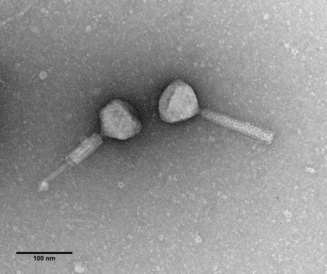
MSPA55 bacteriophages isolated from P. aeruginosa with long tails, which can be seen here in the contracted and non-contracted forms (Maria Stroyakovski, Prof. George PC Salmond group, Biochemistry, Cambridge).
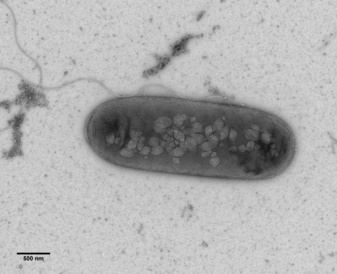
Bacterium with flagellum and gas vesicles (Rita Monson, Prof. George PC Salmond group, Biochemistry, Cambridge).

Exosomes isolated from cell culture supernatant: immuno-staining using the exosome marker anti-CD63 and a secondary antibody tagged with 15 nm gold particles (arrows), followed by uranyl acetate staining (Karin Müller, Prof. Catherine Shanahan, King’s College, London).
